| OCR Text |
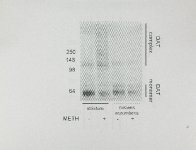
Show 37 80S, 18% glycerol, 180 mM Tris base, pH 6.8, and bromophenol blue) and frozen at -80 CC until Western blot analysis. 10 Equal quantities of total protein (4- IJg) were loaded onto a 4-12% NuPAGE Novex Bis-Tris Midi gradient gel (Invitrogen, Carlsbad, CA) and electrophoresed using a XCell4 Surelock Midi Cell (Invitrogen). Samples were then transferred to a polyvinylidene difluoride transfer membrane hybridization Waltham, MA). (Perkin Elmer Western blot analysis (Hadlock et aI., 2009). was Life and Analytical Sciences, performed as described previously Overall DAT complex immunoreactivity was defined as immunoreactivity greater than -120 kDa and was determined for data presented in Fig 2.1. As the majority of OAT complex immunoreactivity shifts to higher molecular weights at 72 h and remains at 7 d following METH-treatment, only the highest molecular weight regions of Western blots were used for determining OAT immunoreactivity as presented in Figures 2.4 and 2.6. OAT was detected using a rabbit polyclonal N-terminal OAT antibody (generously provided by Dr. Roxanne Vaughan, University of North Dakota). VMAT2 was detected using a rabbit polyclonal antibody (AB1767; Millipore, Billerica, MA). GFAP was detected using a mouse monoclonal antibody (556329; BD Bioscience, San Jose, CA). Data analysis. Analysis of variance, followed by Newman-Keuls post hoc test, was performed to determine significant differences among experimental groups. Differences were considered less than or equal to 5%. All significant if the probability of error was statistical analyses were GraphPad Prism 5 (GraphPad Software Inc., San Diego, CA). performed using |