| OCR Text |
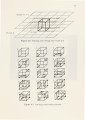
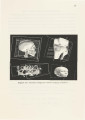
Show 16 The surface intersects those cube edges where one vertex is outside the surface and the other is inside the surface. This means that the data value at one vertex is greater than or equal to the isovalue and the data value at the other vertex is less than the isovalue. • The location of the intersection is found by linear interpolation. • The topology of the surface is obtained from the complementary and rotational symmetry of the 15 patterns illustrated in Ref. [26] and Figure 3. 7.1 • The image is shaded using gouraud shading. Normals are calculated at each voxel using central differences along the three coordinate axes by: G ( . . k) = D(i + 1,j, k)- D(i- 1,j, k) x z, J, /j.x (3.4) G ( .. k) = D(i,j + 1,k)- D(i,j -1,k) y 'l,J, fj.y (3.5) G ( .. k) = D(i,j,k+1)-D(i,j,k-1) z z, J, /j.z (3.6) The gradient at the point of intersection is obtained by linear interpolation. Figure 3.8 shows isosurfaces for the CT data, MRA data, Magnetic Resonance Imaging (MRI) data for knee and brain. The Slicer provides us with the ability to view an arbitrary 2D slice through the volume[5]. The volume can be rotated around any of the axis and any orthogonal slice through this volume can viewed. 1There are well-known problems with the topology given here. It leaves holes in the surface that is generated. This topology will suffice for our purpose of utilizing the slicer /isosurface combination for specifying regions of interest. |