| OCR Text |
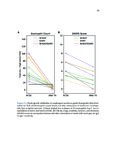
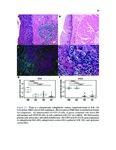
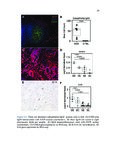
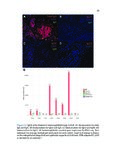
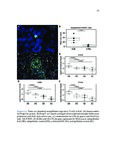
Show 95 B.3 Genomic landscape of metastatic hormone-sensitive vs. castration-refractory prostate cancer B.3.1 Authors Andrew W. Hahn, Edwin Lin, John Esther, Neysi Anderson, Nityam Rathi, Mark Yandell, Benjamin L. Maughan, Neeraj Agarwal B.3.2 Introduction Metastatic castration-refractory prostate cancer (mCRPC) carries a poor prognosis, and targeted therapies have had minimal success in mCRPC. Novel genomic targets could improve drug development. To date, large ctDNA studies in metastatic prostate cancer have been descriptive with limited or no clinical annotation. Herein, we hypothesize that profiles of genomic alterations (GAs) in ctDNA not only differ significantly between, but can also be used to predict mCRPC versus metastatic hormone-sensitive prostate cancer (mHSPC). These findings could help identify new drug targets for mCRPC treatment. B.3.3 Methods Men with mHSPC or mCRPC who underwent NGS of ctDNA using G360 (Guardant Health Inc.) at the Huntsman Cancer Institute were included. Men were classified as mCRPC or mHSPC (patients with current or no prior ADT). G360 detects somatic mutations in selected exons of 73 genes, amplifications in 18 genes, and selected fusions in six genes. Two-sided students t-test was used to compare the %cfDNA and total GAs. The Chi squared test was used to compare the frequency of each GA. Machine learning (ML) algorithms were trained on GAs and benchmarked by cross-validated performance. GAs contributing to mCRPC versus mHSPC classification were measured by ML feature importance (e.g., odds ratios, regression coefficients). B.3.4 Results Of the 259 men included, 119 men had mHSPC and 140 had mCRPC. Men with mCRPC had more GAs (4.5 vs. 1.86, p<0.0001) and higher %cfDNA (9.56% vs. 5.02%, p=0.02). In mHSPC, there was no significant difference in the number of GAs or %cfDNA between men on ADT and those who had not yet started ADT. ML algorithms used GAs to predict mCRPC with 78.1% sensitivity, 64.0% specificity, 76.7% PPV, 65.1% NPV, and 70.3% |