| OCR Text |
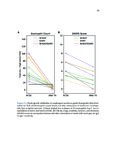
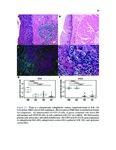
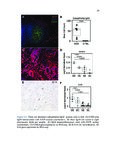
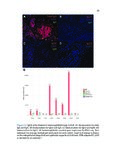
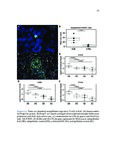
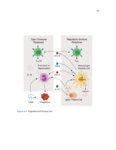
Show 42 invasive are needed. Recent studies have used RNA-sequence analysis and machine learning to create EoE diagnostic tools and identify specific EoE phenotypes [4]. Herein, we present data on high-throughput discovery of candidate EoE genes using machine learning algorithms, where we specifically targeted EoE genes with protein transcription products. Esophageal secretions can be reliably obtained with the patient awake, at substantially lower cost and risk to the patient compared to endoscopy [5]. We assessed whether expressed proteins could be collected in esophageal luminal secretions and used to differentiate between patients with active EoE, treatment-resolved EoE, and controls. 4.2 4.2.1 Methods Potential biomarker identification via transcriptome analysis In order to discover candidate biomarkers for EoE in a high-throughput manner, we analyzed RNA-sequence data from a previous study consisting of 14 EoE patients and 14 controls [6]. Decision trees, a type of machine learning algorithm, were fitted to gene expression values, measured in transcripts-per-million reads (TPM). Five-fold cross validation was used to assess the diagnostic accuracy of each gene. Genes were designated as candidate biomarkers if the following criteria was met: diagnostic accuracy was 100%, and the gene encodes a secreted protein product that can be measured using a commerciallyavailable assay. 4.2.2 Biomarker validation in esophageal secretions The diagnostic accuracy of candidate biomarkers was assessed at the protein level in esophageal secretions. Esophageal secretions from five patients with active EoE (EoE), six with patients with treatment-resolved EoE (RES), and six controls (CTRL) were obtained using an endoscopic cytobrush. For each sample, protein expression was assayed using a Luminex 23-cytokine panel. 4.3 Results Genes encoding potential protein biomarkers were identified via transcriptome analysis, including CCL26, which encodes eotaxin-3, KITLG, which encodes stem cell factor (SCF), and POSTN, which encodes periostin (Figure 4.1A-C). Eotaxin-3 was highly ex- |