| OCR Text |
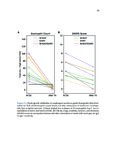
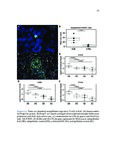
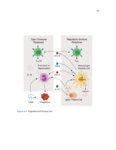
Show 36 PETERSON | ET AL. T A B L E 1 Clinical characteristics of EoE and PPI-REE patients DE genes in FFPE vs RNA–later (A) EoE (N = 14) RNA–later Log2 FC, P–adj < 0·01 6 3 0 –3 38 (22-62) 48 (24-80) 43 (18-75) 71% 69% 64% Max.tissue eos./hpf (mean, range) 39 (16-65) 35 (22-65) Seasonal allergic conjunctivitis 62% 54% Asthma 25% 15% Allergic rhinitis 57% 54% 7% 18% IgE (IU/mL serum) (B) –3 0 3 FFPE Log2 FC, P–adj < 0·01 6 DE genes in FFPE vs Sherrill et al Sherrill et al Log2 FC, P–adj < 0.01 6 Controls (N = 14) % Male Allergic eye symptoms –6 PPI-REE (N = 13) Age (years, range) Atopic dermatitis –6 3 57% 399 0 (0-0) 54% 542 The mean age and gender of all 3 groups (EoE, PPI-REE and normal controls) did not significantly differ. Additionally, the maximal oesophageal tissue eosinophil content, IgE and clinical comorbidities did not differ significantly. Statistical comparison was done by Mann-Whitney U test for age, eosinophil counts and serum IgE. Proportions were compared by Fisher’s exact test. P = 0.20 for age (EoE vs PPI-REE). For all other comparisons, P ≥ 0.37. EoE, eosinophilic oesophagitis; PPI-REE, proton pump inhibitor-responsive oesophageal eosinophilia 3 2.3 | Formalin-fixed, paraffin-embedded data set validation 0 While RNA yields from formalin-fixed, paraffin-embedded tissues are similar to those of RNAlater-preserved tissue,23 RNA extracted from –3 formalin-fixed tissue is degraded to smaller fragments (typically peaking at 200-300 bases) than RNA from RNAlater-preserved tissue. Our validation study compared gene expression using formalin-fixed, –6 paraffin-embedded tissue vs RNAlater-preserved samples. We used –6 –3 0 3 FFPE Log2 FC, P–adj < 0·01 6 F I G U R E 1 Validation comparing EoE/control expression ratios— formalin-fixed, paraffin-embedded tissues vs RNAlater, and formalinfixed tissues vs a prior study. To validate the ratios of disease to normal RNA extracted from formalin-fixed tissues, we compared the EoE to control log 2 ratios for formalin-fixed, paraffin-embedded extracted RNA to those of well-preserved RNA. Only genes differentially expressed (DE) at the adjusted p/FDR < 0.01 level are used. In each, the formalin-fixed tissue results are on the x axis and well-preserved RNA results on the y axis. A, In a separate, preliminary data set, we compared our EoE formalin-fixed tissue results (n = 7 EoE and 6 controls) vs results from an RNAlater group (n = 6 EoE and 6 controls). There were 1529 differentially expressed genes, with excellent correlation; Pearson product moment correlation r = .96. B, We also compared our full formalin-fixed, paraffin-embedded tissue results (n = 14 EoE and 14 controls) to those of Sherrill et al24 (n = 10 EoE and 6 controls). There were 624 differentially expressed genes, with very good correlation; Pearson product moment correlation r = .92. Both results are highly significantly correlated (P < 2.2 9 10 16 for each). EoE, eosinophilic oesophagitis; FDR, false discovery rate Illumina sequencing chemistry based on 50-base pair reads—indicating formalin-fixed tissue would be a viable alternative. We compared 7 formalin-fixed EoE oesophageal biopsies to 6 formalin-fixed controls, and 6 EoE RNAlater-preserved biopsies to 6 RNAlater-preserved normal controls. Our full study formalin-fixed, paraffinembedded EoE and control data sets were similarly compared to those of Sherrill et al (Figure 1B)24 which also used RNAlater-preserved mRNA. 2.4 | Real-time quantitative reverse transcription PCR validation Subtle biological changes could be missed by DESeq2 statistics, so we also examined genes with 2-fold differences that were significant at P < 0.05. Thirty of our 41 RNA-Seq-analysed formalinfixed RNA samples were used in Real-Time Quantitative Reverse Transcription PCR; 11 samples had an insufficient RNA after their use for RNA sequencing. Consequently, our real-time quantitative reverse transcription PCR sample set constituted these principal component analysis were performed using Cluster 3.0 and numbers per group: 12 EoE, 11 PPI-REE and 7 Control. For the R application “rgl”, respectively, to identify unique and common these 30 RNAs, total RNA was reverse transcribed to cDNA patterns of expression across each sample. using the Applied Biosystems High Capacity RNA-to-cDNA Kit. |