| OCR Text |
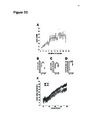
Show 68 TABLE 4.2 | Experimental numbers Group Control LPS 6hr LPS 24hr Animals Slices 6 4 4 20 11 16 Cells ROIs ROIs/Cell 135 106 105 480729 395493 358545 3561±756 3731±724 3415±719 Events (Process) 24849 24313 7904 Events (Soma) 45 29 11 tdTomato channel, was presented to the user who was then prompted to select the soma, cell boundaries and areas to exclude such as neuronal cell bodies. tdTomato robustly labels fine processes, so selection of the cell boundaries was usually straightforward. The selected somatic area was treated as a single ROI. Process ROIs were assigned by splitting the area overlying the processes into 1 µm2 ROIs (Figure 4.1B). Because protoplasmic astrocyte domains are densely packed with fine processes, little dead-space was included in ROIs. A raw fluorescence intensity time series (F) was then generated for each ROI by averaging the constituent pixel values at each time point (Figure 4.1C) and was then converted to (F - F0)/F0 (ΔF/F0) with F0 set to the median of F. An amplitude threshold was used to identify suspected events. All points with a ΔF/F0 greater than 2.5 times the interquartile range above the third quartile (Q3+2.5*IQR) for each individual ROI trace were considered to belong to a suspected event. Each suspected event, with a peak amplitude of ΔF (Figure 4.1D left), was fitted to a (symmetric) Gaussian with a peak equal to ΔF and the same full width at half-maximum (FWHM; Figure 4.1D middle). The integral of the event ΔF/F0 with respect to time was then approximated as the area under the curve (AUC) of the Gaussian (Figure 4.1D top right). An additional thresholding was applied to process events. Due to the low baseline fluorescence in processes, small fluctuations in intensity caused by shot noise can result in noise activity above threshold. These tend to have a duration of one frame period, only include a single point above threshold and are not |