| OCR Text |
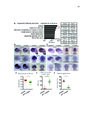
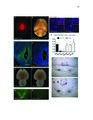
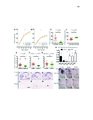
Show 39 STEM CELLS AND REGENERATION Development (2016) 143, 45-53 doi:10.1242/dev.126813 regeneration may require these divisions at an even higher frequency. However, our data indicate that ectopic Wnt activity in fact inhibits expansion of the hypothalamic radial glia population, while other studies suggest that signals such as FGF (Kaslin et al., 2009; Robins et al., 2013) and Sonic hedgehog (Dave et al., 2011; Shikata et al., 2011; Komada, 2012) are likely to promote this process. We found that a specific subset of lef1-dependent neurons located near the hypothalamic ventricle arise from the radial glial lineage (Wang et al., 2012). Combined with other studies in the retina (Agathocleous et al., 2009), cerebral cortex (Munji et al., 2011; Zhang et al., 2014), hippocampus (Seib et al., 2013) and midbrain (Castelo-Branco et al., 2003), our data suggest that the most widely conserved role for Wnt/β-catenin signaling in the CNS might be to regulate the differentiation of specific subsets of committed neural progenitors. Wnt/β-catenin activity does not act identically in all neural stem and progenitor cells Our experiments support the idea that diverse neural stem and progenitor cell populations are likely to exhibit different responses to Wnt/β-catenin signaling. Although Wnt ligands and reporters are expressed at high levels in the hypothalamic ventricular zone (Wang et al., 2012), radial glia largely fail to respond to these signals. This low activity state could be regulated by extracellular or intracellular pathway antagonists, or radial glia might simply fail to express the appropriate receptors to transduce Wnt signals. Regardless of the mechanism, it appears that this characteristic of hypothalamic radial glia is similar to other radial glial populations in the zebrafish retina and spinal cord (Goldman, 2014; Briona et al., 2015), but differs from radial glia in the mammalian dentate gyrus (Qu et al., 2010). Other studies have similarly shown that neural progenitor populations vary dramatically in their interpretation of pathway activity (Poschl et al., 2013). Understanding these differences might help provide insight into the basis of radial glial, and neural stem/ progenitor cell, heterogeneity. MATERIALS AND METHODS Fig. 6. Wnt/β-catenin signaling activation blocks expansion of the radial glial population. (A) Induction of Wnt8a at 5 dpf leads to a decrease in the number of GS+ radial glia at 6 dpf. (B) Addition of 4 µM BIO, a Gsk3β inhibitor, daily from 6-9 dpf leads to a decrease in GS+ radial glia. (C) Representative images of labeling for GS in control and BIO-treated larvae. Images are single optical sections from ventral views of whole-mount brains. Scale bar: 10 µm. (D) Following recombination from 5-6 dpf, incubation in BIO causes a significant decrease in the number of labeled progeny at 9 dpf. (E) Incubation in BIO causes a significant increase in the percentage of GS+ cells within the labeled lineage. Error bars indicate s.e.m.; n=40 optical sections from four brains for each experiment. that Wnt/β-catenin signaling might function generally to promote stem and progenitor cell proliferation. In order to achieve the increase in population size that we observe in our lineage analysis, radial glia must undergo amplifying self-renewing divisions, and, as has been shown in the telencephalon (Barbosa et al., 2015), Embryos were obtained from the following zebrafish lines: Tg(her4.3: EGFP)y83 (Yeo et al., 2007), Tg(ubi:loxP-eGFP-loxP-mCherry)cz1701 (Mosimann et al., 2011), Tg(−3her4.1:ERT2-Cre-ERT2)vu298 (Boniface et al., 2009; Mosimann et al., 2011), Et(Gal4-VP16,myl7:gfp)zc1066a (Wang et al., 2012), Tg(UAS-E1b:NTR-mCherry)jh17 (Davison et al., 2007; Pisharath et al., 2007), Tg(7xTCF-Xla.Siam:GFP)ia4 (Moro et al., 2012), Tg(hsp701:dkk1-GFP)w32 (Stoick-Cooper et al., 2007), Tg(hsp70l:wnt8aGFP)w34 (Weidinger et al., 2005), lef1zd11 (Wang et al., 2012), Tg(th2: GFP-aequorin)zd201 and Tg(th2:Gal-VP16)zd202 (McPherson et al., 2016). Embryos were staged according to Kimmel et al. (1995). All experiments were approved by the University of Utah Institutional Animal Care and Use Committee. Transgenic embryos were identified by GFP tag expression following heat shock induction of wnt8 or dkk1, or by PCR amplification of trunk tissue for dkk1 induction in the presence of the -3.5ubi:loxP-EGFP-loxPmCherry reporter using the following primers (5′-3′): dkk1 forward, TCGACTCAAGGATCACCACA; gfp reverse, TCCCTCAAACTTGACTTCAGC. lef1 mutant animals were identified by the absence of posterior neuromasts as labeled with DASPEI (Invitrogen) (McGraw et al., 2011; Wang et al., 2012). Treatment of embryos and larvae Cre-mediated recombination was performed by incubation in 5 µM 4-hydroxytamoxifen (4-OHT; Sigma, CAS RN 68047-06-3) in 1% DMSO from 5-6 dpf. Ablations were performed by incubation in 1 mM 51 DEVELOPMENT Zebrafish |