| OCR Text |
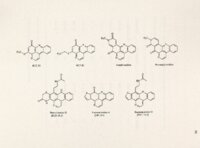
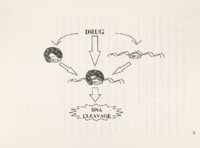
Show 83 mechanisms of drug-enzyme interaction. Cleavage site specificity was unknown for ascididemin and its analogs, but if it were' dependent on nucleotide specific interactions, a DNA cleavage pattern might emerge. the Non-specific DNA fragmentation is distinct from site-specific cleavage pattern of topoisomerase-targeted drugs. Mitra et al., 1997, reported nonspecific DNA cleavage by leinamycin, a natural product antibiotic, through a thiol-dependent, oxidative mechanism. Multiple, frequent, nonspecific DNA fragmentation observed with all three DNA active compounds suggested a similar nonspecific mechanism of DNA cleavage among ascididemin, BC-1 09-1 and BCMH-1-48. This DNA cleavage pattern was different from the cleavable inhibitors. complex formation at preferred sites seen with classical topoisomerase Although Bonnard et al., 1994 reported DNA intercalation of ascididemin with a binding preference for GC-rich sequences, our results did not show a preference of ascididemin cleavage for particular nucleotides or DNA sequences in the presence of DTT. A non-specific pattern was consistent with an ROS mechanism since ROS have been reported to react the indiscriminately with closest neighboring molecules (Schraufstatter et al., 1988). It is also known that reactive oxygen species can abstract H+ from the deoxyribose sugars of either purine or pyrimidine bases and lead to formation of single strand breaks (Colton and Zakhari, 1997). The ethidium bromide displacement assay revealed likely DNA intercalation of ascididemin, BC-109-1, BCMH-1-48, BC-1-21 and BC-1-31. make them good candidates for DNA intercalation. Their planar structures Other marine pyridoacridines (i.e., dercitin, cystodytin J and diplamine) were previously shown to intercalate into DNA |